How
to use ReConn?
Retrieving pathway information:
There are three main ways to retrieve pathway information using ReConn:
Search for a pathway by its name
Grow a pathway by starting with a certain
compound
View all reactions
Mapping of data
Data can be mapped using the two following ways:
Expression colouring
PathwayFinder
Colouring nodes by occorunce
Occurence colouring
Various options of interacting with the network in ReConn
Expanding pathway nodes
Retrieving hypothetical neighbours
Calculating alternative
routes
Finally one could use a locally installed Reactome database.
Using a local Reactome database
Search for a
pathway by its name:
* Select Query database from ReConn in the Plugin menu:
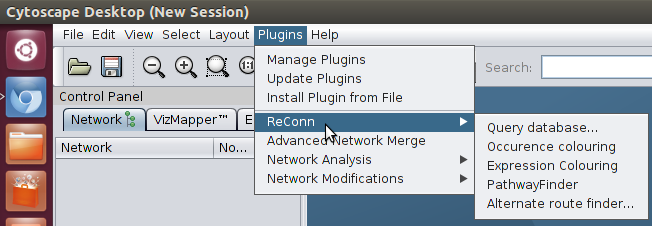
* The Query options dialog appears. Here you can specify which data you
want to get from Reactome.

* For instance only get human reactions:
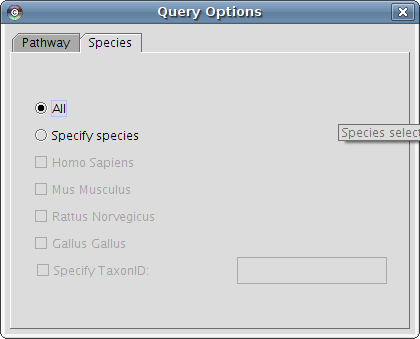

* Then return to the first tab and enter (a part of) the name of the
pathway.

* The result:
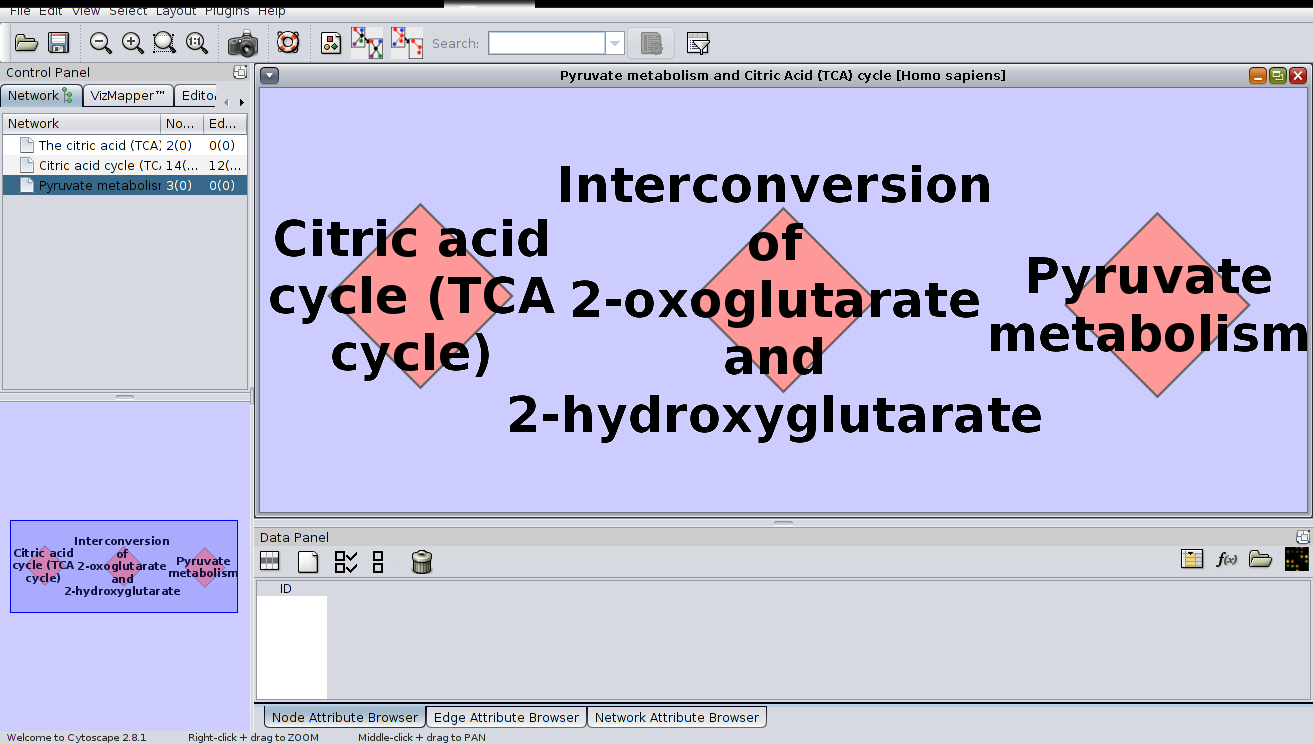
Selecting the second network:

The
circles are reactions, the diamond shapes are pathways and rounded squares are BlackBoxEvents. The edges
respresent (one or more) metabolites passing between reactions. The
labels in the node are the genes associated with the reaction. The
mouse-over gives you the reaction information. Or if you hover over an
edge, it gives you the metabolite that is shared between the reactions.
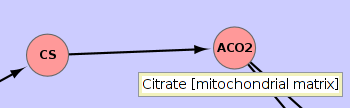
Grow a pathway by
starting with a certain compound
Besides
selecting a certain pathway one can also "grow" a pathway. You just
have to specify the metabolite from which to start. Again only human
reactions are chosen.
A valid search term is a case insensitive full name of a compound as it is stored in Reactome.

* result:

Here we have rearranged the pathway nodes for readability.
The rounded square nodes, represent BlackBoxEvents.
View all
reactions
One can also select all reactions for visualization at once. Again only
human reactions are chosen.

*result:

Expression
colouring
Expression data can also be used to colour the nodes:


Here
the tab delimited file can be specified that should be used. The first
column should contain the ID (e.g. AffyMetrix id) to be used for the mapping. And the column
that should be used for colouring is specified with the drop down list.
Example expression colouring file
One can also specify conditions that should be met, for
colouring.

result of Expression colouring on the pathway grown from insulin.
PathwayFinder
Instead
of getting a pathway and visualize data on top of it, one can also use
the data to find the pathways that are of interest.

Similar as to Expression colouring, ABS is used to specify if the
absolute value of the data should be used for the condition.

A
new window pops up (after a while). For each pathway the amount of
reactions meeting the criteria as well as the total number of reactions
are specified.
Select a pathway and click on "Show pathway" to visualize the data on
top of that pathway.
Occurence colouring
The occurence of a reaction in different species can be used to colour
the nodes. A whiter node occurs in less species.


Expanding
pathway nodes
One can open a pathway node by right clicking on the diamond shaped
node, to get the contextmenu:

* and select Open Pathway. All reactions in the selected pathway are
now added to the current network.

Retrieving hypothetical neighbours
To get the hypothetical neighbours of a node or nodes, select the nodes
and right click on them to get the contextmenu.

* result:
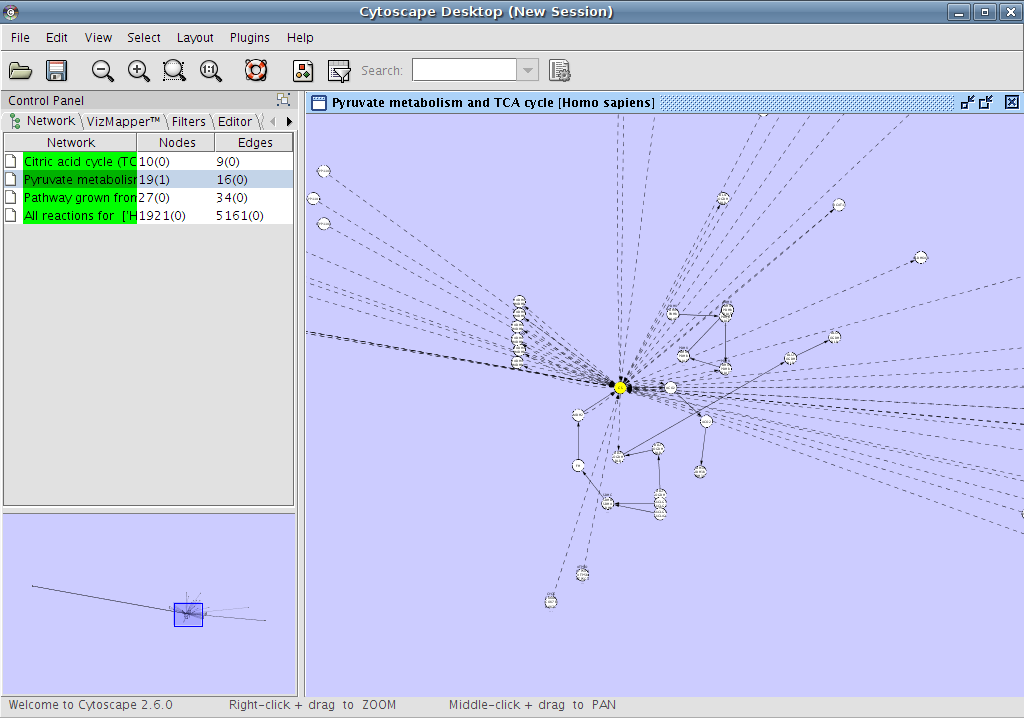
The hypothetical neighbours have dashed lines.
Calculating alternative
routes
This
feature calculates the subgraph containing all routes from one reaction
another. The user can specify the maximum pathlength as well as
reactions that are not allowed. (For instance reactions that are
knocked out)
The most simple way of using this feature is by specifying the "start",
"end" and "not using" nodes by using the context menu.
Just
select the reaction(s) you want to be the start and select "Start
reaction" from the context menu. Do the same for the end reaction(s)
and the ones you don't want to use.
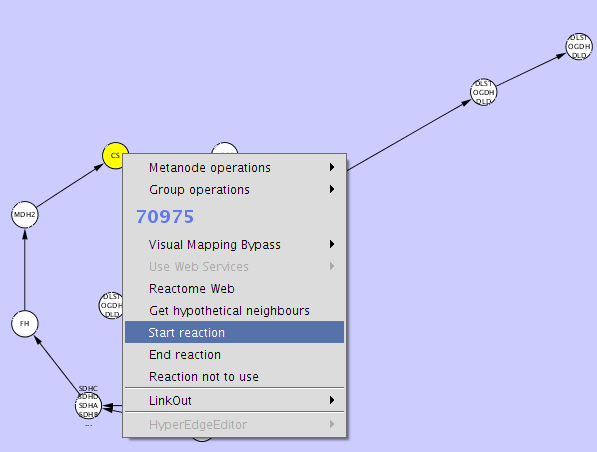
After
you have done this, you can select the Alternate route finder from the
ReConn menu. You will see that most of the field have already been
filled out.
You will only need to specify the depth of the search.


This results in:

You can also enter the values of start, end and not using
fields manually by specifying their Reactome ID. It is
possible to have multiple start and end points, as well as multiple
reactions that should not be used. The numbers should then be seperated
by a ;.
The Reactome Id can be found on the Reactome website by looking at the
URL.
http://www.reactome.org/cgi-bin/eventbrowser?DB=gk_current&ID=71406&ZOOM=2
The ID is after ID=. So in this case it is: 71406
Using a local Reactome
database
A local instance of Reactome can also be used. The MySQL dumps can be
found here: sql
ReConn also needs the denormalized database found here: sql_dn
The
set this up, the use should put a tab-delimited text file called
server.tsv in the Cytoscape main directory NOT THE PLUGIN directory.
The file should look have the following layout:
server<tab>databasename<tab>user<tab>password<tab>port
server<tab>databasenameDN<tab>user<tab>password<tab>port
example:
bmi.bmt.tue.nl\treactome\tguest\t\t3306
bmi.bmt.tue.nl\treactome_dn\tguest\t\t3306


























